Regulation of simultaneous production of useful proteins in filamentous fungi


Expression Regulation Network Construction Technology
Nagaoka University of Technology, Kao Corporation, Japan Bioindustry Association, Kyushu University, and National Institute of Advanced Industrial Science and Technology (AIST)
To establish a technology platform for constructing strains producing optimal enzyme preparations for saccharification of plant biomass of various components, by controlling the production ratio of each plant biomass saccharifying enzyme excreted by filamentous fungi.
Plant biomass is a sustainable source of energy, existing abundantly on the earth. The sugar obtained by hydrolyzing (saccharifying) the polysaccharides that make up the plant cell wall, which is a non-edible part of plants, is not only used as a raw material for biofuels, but also usable as a carbon source for the production of high value-added products in microorganisms. The technology for saccharifying plant cell walls is essential also for creating a bioeconomy society based on material production using microorganisms. Since filamentous fungi produce an enzyme "cellulase" derived from a microorganism capable of efficiently saccharifying plant cell wall polysaccharides, research and development aiming at enzymatic saccharification of plant biomass using cellulase are underway. In Japan, development using a filamentous fungus Trichoderma reesei, which has a high cellulase productivity, has been promoted since the 1980s. Initially, capability to produce enzymes was improved by inducing mutations, and the finally acquired PC-3-7 strain is still used as the base strain for the development of industrial enzyme production strains. In addition, the production of saccharifying enzyme-producing strains that have the world's top level plant biomass saccharification capacity was succeeded through the production of an enzyme that plays an important role in saccharification in the PC-3-7 strain using genetic recombination technology. However, since the composition of the constituents of the cell wall polysaccharide differs depending on the type of plant biomass, the creation of a "tailor-made saccharifying enzyme," which consists of the optimal enzyme components for the plant biomass, is currently being required (Fig. 1).
Strain developments so far have been based on previously published papers and findings obtained in the laboratory. In such developments, although it was possible to control the total production level of enzymes, controlling only the production levels of specific enzymes was difficult. However, the identification of factors that control multiple genes is difficult with the conventional "wet" approach that relies on the experience and intuition of researchers. Therefore, in the Smartcell Project, we aim to clarify which gene regulates which gene and how the network of regulation is structured and also to identify the factors regulating the production ratio in the group of glycoside hydrolases, by preparing a large amount of expression data on all the T. reesei genes and making the best use of the network model construction technology of the smart cell design system.
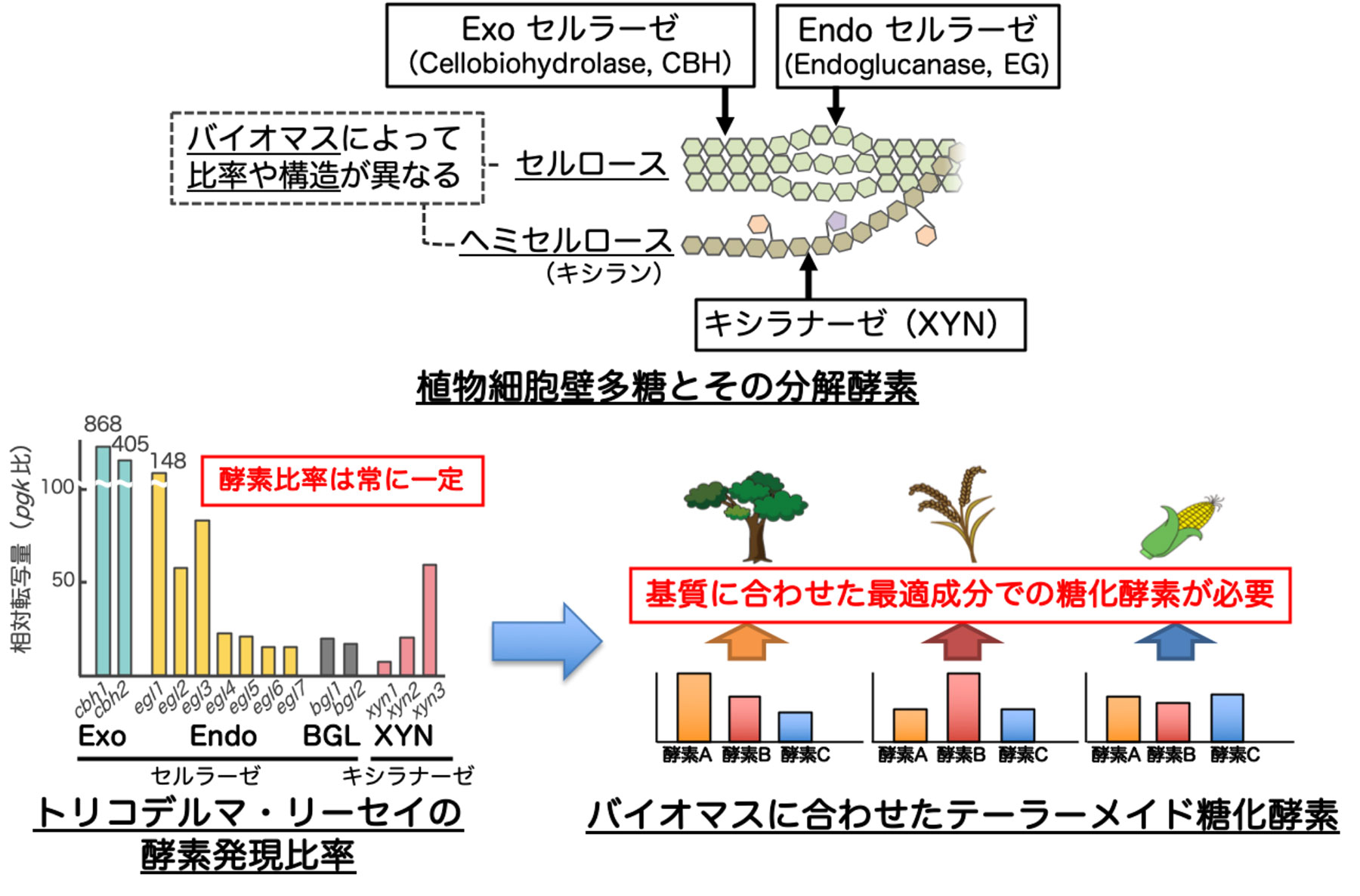
図1 本研究開発の背景
Using approximately 200 samples of all gene expression data under the cellulase production and non-production conditions, we designed a gene expression network based on the DBTL cycle shown in Fig. 2, and estimated multiple genes that may affect the gene expression of the target enzyme. By constructing knockout strains of these genes, we analyzed the cellulase productivity and the enzyme production ratio. As a result, we were able to find regulatory genes that control the production of specific cellulases. By utilizing these regulatory genes, we have succeeded in improving the saccharification capacity for a specific plant biomass. Although the constructed gene control network is a prototype, we are pursuing information analyses further to improve accuracy based on feedback from our achievements obtained so far. However, even at present, it has been found that genes whose existence was extremely difficult for researchers of T. reesei to predict significantly affect the production balance of enzyme components. Therefore, we expect to generate new findings by further promoting this research.
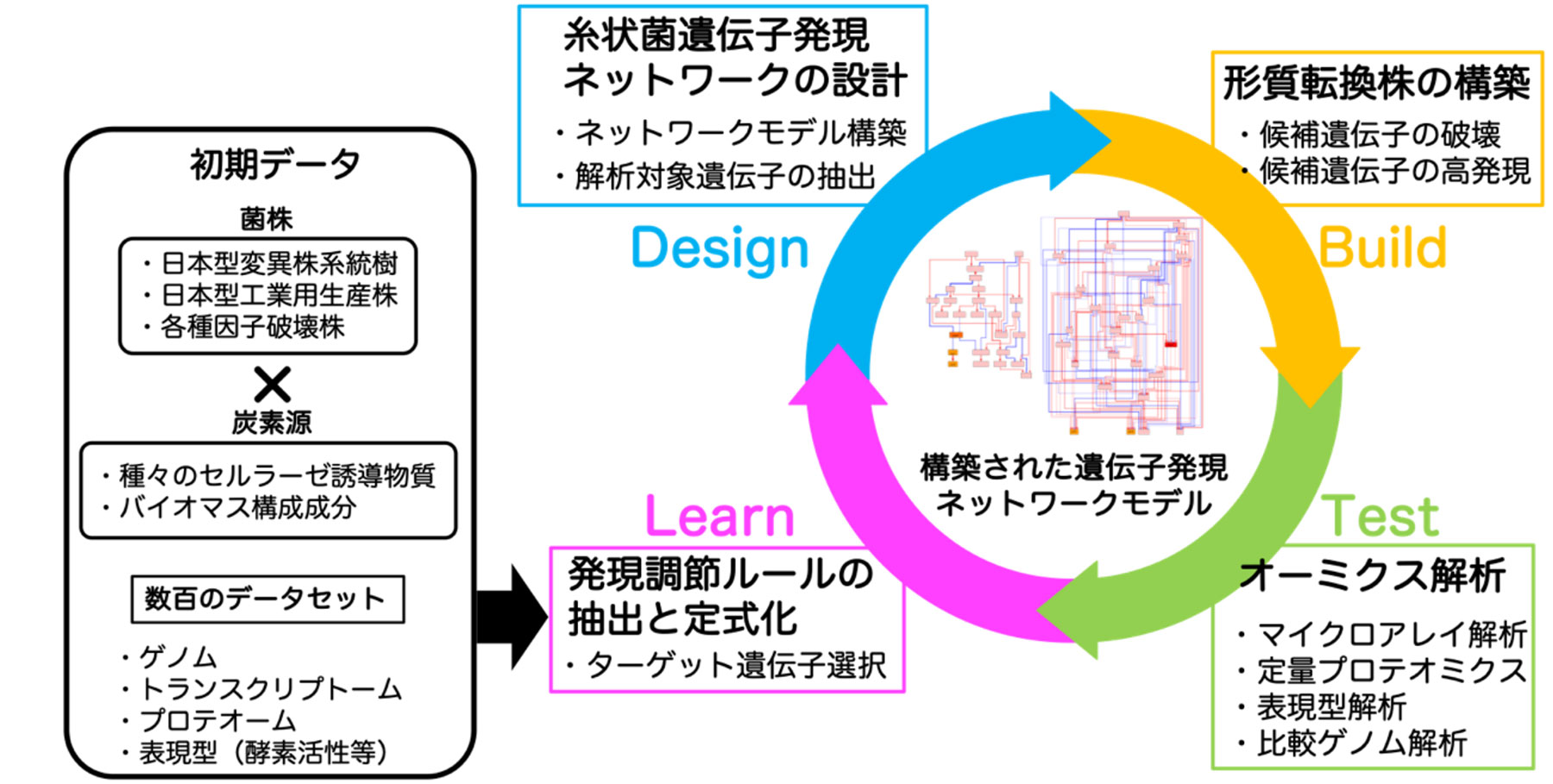
図2 本研究開発の流れ
Last updated:December 25, 2023